SageDDA, SageDIA and SagePRM analysis
After an experiment has been created, and raw spectral files have been uploaded to the experiment, SageDDA, SageDIA or SagePRM analysis can be selected.
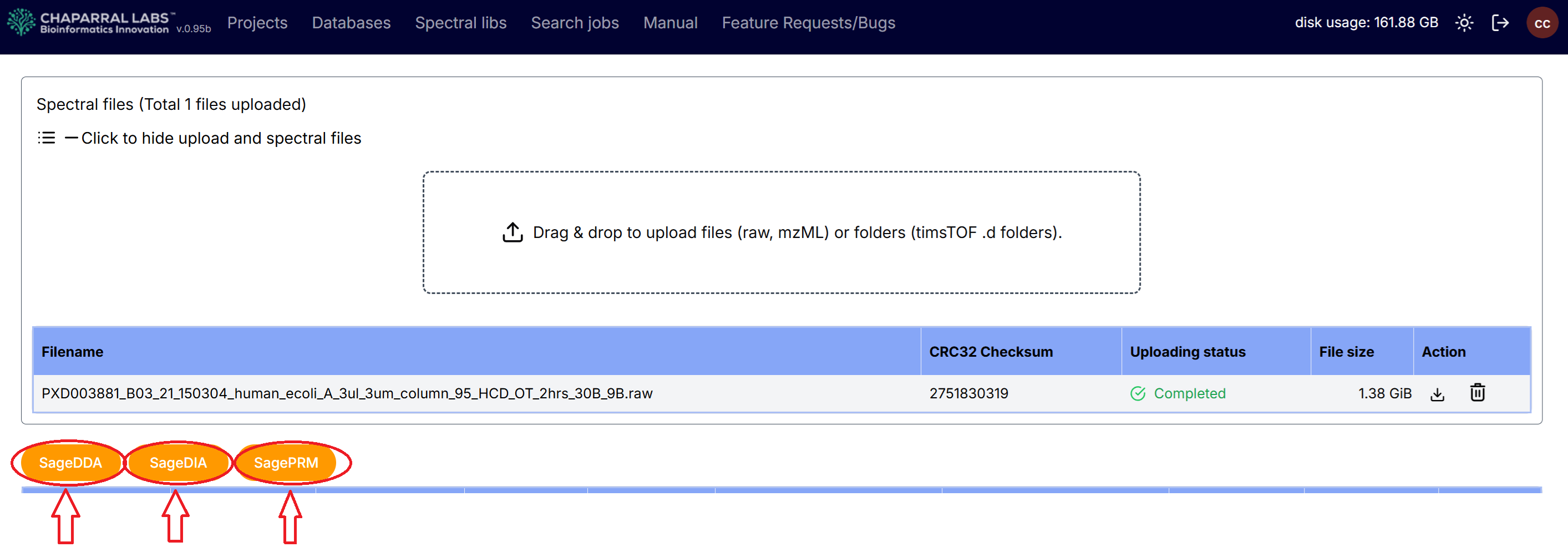
After an experiment has been created, and raw spectral files have been uploaded to the experiment, SageDDA, SageDIA or SagePRM analysis can be selected.
